Visualizing White Matter Structure of the Brain using Dijkstra’s Algorithm
-
. Visualizing White Matter Structure of the Brain using Dijkstra’s Algorithm. In Peter Zinterhof, Sven Lončarić, Andreas Uhl, and Alberto Carini, editors, Proc. 6th International Symposium on Image and Signal Processing and Analysis (ISPA 2009, September 16--18, Salzburg, Austria), pages 575--580, 2009.
Download: pdf
Abstract
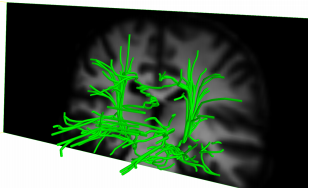 An undirected weighted graph may be constructed from diffusion weighted magnetic resonance imaging data. Every node represents a voxel and the edge weights between nodes represent the white matter connectivity between neighboring voxels. In this paper we propose and test a new method for calculating trajectories of fiber bundles in the brain by applying Dijkstra's shortest path algorithm to the weighted graph. Subsequently, the resulting tree structure is pruned, showing the main white matter structures of the brain. The time consumption of this method is in the order of seconds.
An undirected weighted graph may be constructed from diffusion weighted magnetic resonance imaging data. Every node represents a voxel and the edge weights between nodes represent the white matter connectivity between neighboring voxels. In this paper we propose and test a new method for calculating trajectories of fiber bundles in the brain by applying Dijkstra's shortest path algorithm to the weighted graph. Subsequently, the resulting tree structure is pruned, showing the main white matter structures of the brain. The time consumption of this method is in the order of seconds.